












• •
出版日期:2026-03-03
通讯作者:
赵莹(1982—),女,博士,副研究员,研究方向:实验动物遗传。E-mail: zhaoying@slarc.org.cn,ORCID: 0000-0002-0551-0566作者简介:汤建平(1974—),本科,实验师。研究方向:实验动物生产管理。E-mail: tangjianping@slarc.org.cn
TANG Jianping1,2( ), ZHAO Liya2, ZHAO Ying1(
), ZHAO Liya2, ZHAO Ying1( )(
)( )
)
Published:2026-03-03
Contact:
ZHAO YING(ORCID:0000-0002-0551-0566), E-mail: zhaoying@slarc.org.cn摘要:
目的 筛选一套覆盖大鼠1-20号常染色体和X染色体、且每条染色体含2-4个的STR(Short Tandem Repeats, STRs)标记,建立适用于F344、BN、DA、Lewis、PVG这5个常用近交系大鼠的遗传背景检测、遗传污染检测及品系间鉴定的专用标记组合。 方法 每个品系抽样6只,雌雄各半,抽提基因组DNA。从国标、文献中分别选取27个和34个大鼠STR标记,共计61个。采用单重STR PCR扩增结合毛细管电泳技术对5个近交系大鼠样本进行分型,利用GeneMapper ID v3.2软件分析STR基因型数据。基于STR分型结果,通过GenAlEx 6.51b2软件计算品系间遗传距离,并借助MEGA7软件构建近交系大鼠遗传进化树。 结果 在61个STR标记中,D15mit3存在非特异扩增,D3wox7无特异性扩增产物,最终可用的STR标记共计59个。该59个STR标记在5个近交系大鼠品系中特异性扩增成功率达100%,且每个样本的分型结果均为纯合,5个近交系大鼠的平均纯合度达100%,符合近交系遗传质量控制的要求。遗传进化树显示,BN与F344的STR标记等位基因差异数为49,遗传距离达1.775,为所有品系对中的最大值,表明二者遗传分化程度最高、亲缘关系最远;F344、DA、PVG品系聚为一支,表明三者起源于同一祖先;BN和Lewis品系聚为一支,表明二者起源于同一祖先,5个品系的遗传亲缘关系与既往报道基本一致。在59个STR分型结果中,LCA、AGT、D5Hmgc2在5个品系间无多态性,其余 56个STR标记具有品系间多态性,且覆盖大鼠20条常染色体和X染色体(每条染色体有2-4个STR标记),因此可作为一套近交系大鼠遗传背景检测的STR标记组合。从56个STR标记中选取42个STR标记可用于近交系大鼠遗传污染检测。D7wox14、D15rat123、D20wox3在5个品系中的扩增产物长度存在品系间差异,可用于5个常用近交系大鼠品系间快速鉴定。 结论 本研究成功筛选出了一套可用于5个常用近交系大鼠的遗传检测STR标记,并建立了分别用于遗传污染检测与品系间鉴定的专用标记组合。
中图分类号:
汤建平,赵丽亚,赵莹.常用近交系大鼠微卫星遗传标记筛选及分析[J]. 实验动物与比较医学.. DOI: 10.12300/j.issn.1674-5817.2025.109.
TANG Jianping,ZHAO Liya,ZHAO Ying. Screening and Analysis of Microsatellite Genetic Markers in Commonly Used Inbred Rat Strains[J]. Laboratory Animal and Comparative Medicine. DOI: 10.12300/j.issn.1674-5817.2025.109.
| 标记Marker | 染色体Chromosome | 物理位置/bp# Physical position/bp | 正向引物序列(5’→3‘)Upstream primer sequence(5’→3‘) | 反向引物序列(5’→3‘)Downstream primer sequence(5’→3‘) | 退火温度/℃ Annealing temperature/℃ |
|---|---|---|---|---|---|
| D1wox18 | chr1 | 100,133,276 | TCACTCTCATTAAACTAGGAATGC | ACTGTTGGGTAACAAAGTTATGG | 53 |
| D1rat345* | chr1 | 112,693,166 | TTCACACAAATGCCACCAGT | CAAAGAGATAGGGCGACAGG | 59 |
| D1wox21 | chr1 | 198,655,873 | GACATGGTGAGTGAAGACCC | TGAGGCTATCCTGAGCTAACTC | 59 |
| D1mgh14* | chr1 | 278,228,767 | CCGCACTGAGCTCTCAGAG | CCCAACCATTGAGCTAGTAAGG | 61 |
| D2rat3 | chr2 | 8,579,280 | TCTCTGGTGGATACCTCACAAA | ACATGATGTGGAATTCCCCC | 59 |
| D2mgh26* | chr2 | 105,075,219 | ATTGGTGATCAGATTTTTTTCTCC | CAGTTGAAAATTCCTGTATCTGG | 57 |
| D2wox15* | chr2 | 188,450,000 | GGTGCTAGTAGACAATAAGATAGAT | TTCATGAGTTTTCACTGTTTGC | 53 |
| D2Arb20 | chr2 | 200,585,161 | ACTAGAGTCAGGAGGCAAAACC | ACCCTATGTGAAATGGTGATGC | 59 |
| D3wox9* | chr3 | 51,821,710 | TTCAGGTATGATTCGGGAAC | ATTGGCGATATCATTTCACTAAC | 59 |
| RH94469 | chr3 | 93,380,189 | CTTATGTTACCTCACAGCCTGG | GGGTGGGCCATCTTTATAATC | 61 |
| D4wox27* | chr4 | 3,044,017 | GGCCTTCATTTGTCCTTTGA | CACCATTATGCCCAGCCTTA | 59 |
| D4Arb10* | chr4 | 61,708,051 | ACTCATCTTCTGAGGAGTAGCG | TTCACAGTCTATTTGGTAGGGC | 59 |
| D4mit15* | chr4 | 111,809,356 | CCTAATGACAGCAATAGAAACTACG | TTTTGGTACTGGAGAAAGTTGG | 61 |
| D5mit18 | chr5 | 17,062,312 | GCAAGACAGGCTTCCACAAT | AATACTGCCGGAGCAGAAAA | 59 |
| D5Hmgc2* | chr5 | 78,041,475 | ATGTTCCCTGATGTCCCTTC | AAACCATGCCACGTGAAC | 53 |
| D5wox11 | chr5 | 138,180,823 | CTTAGGTCAGCTGGTCCCTAG | CATGGTGCTTTTCCTTGTTG | 55 |
| D6rat105 | chr6 | 19,888,103 | GCATGAGGCAGAGCCAATTA | GCCAGTTTGCCTTCACTTTC | 59 |
| D6mit1* | chr6 | 99,273,921 | TTAGGAGAGAACTGAAAGTTGTCC | ATGGTGCACTATGGTGGTCA | 61 |
| D6wox12 | chr6 | 138,065,848 | CTTCCCTACCTTCTACAACACTAAG | TAAACCAAAGAGCATGATGAAG | 61 |
| D7wox14 | chr7 | 102,588,256 | GGGAGTAAAAGAGTGCATTCC | TACCCCAATCCTGAACCAC | 61 |
| D7wox24 | chr7 | 115,250,056 | AGCCTCAATCTGGCCTAGCTC | TGACTCTTGACAATGGCACA | 59 |
| D7mgh3* | chr7 | 123,632,617 | CGGCATGTGTCTCTGTGTG | TGACTTCTGGTGTCCTCACG | 59 |
| D8wox4* | chr8 | 50,529,563 | GATTTGAAGCGATTGTCCAT | GTCTAGCTGCCCACAGGAG | 57 |
| D8rat14* | chr8 | 108,068,142 | GCCCATACGTTGCATCAAGT | GGCCGGTCTAATTATTTCTTCA | 59 |
| D8wox11 | chr8 | 116,873,553 | AGTGGGAATGCTTAGCCAAA | CAGGCCTTTACACCTTCCTG | 57 |
| D9mit2* | chr9 | 71,771,288 | TGACCAGTTAGCCCTTTCCA | GGGAGCAGGGTTCTACACAG | 59 |
| D9rat110 | chr9 | 102,531,710 | TGGGTGCTTAATTTTGTTTGTTT | GCTCCTAACCCATCCAACTTT | 55 |
| D10rat199 | chr10 | 1,381,054 | TGCCAAATAATGATTGTTCCTTC | CCTCCTCTGGCTTCTTGAGA | 55 |
| D10wox12 | chr10 | 56,220,298 | CGCTGGGAGGAAAGAGAG | AGGCTTGCACTCTGTGTTTG | 59 |
| D10mit12* | chr10 | 94,240,320 | ATGGGAGGGAACAAGTCTTC | GAGGGAGAGAGAAAGAGAGACAG | 61 |
| D11wox3* | chr11 | 60,397,957 | CACTTTGAGGCCATTCTGAA | CCTTCTCTTTGTGAAAAATAAAGTC | 59 |
| D11mgh3* | chr11 | 69,649,708 | GGAGCTGAAATACGAGAGAAATAA | GTCCTGCTGGCTGTGCAT | 59 |
| D11wox2 | chr11 | 80,357,513 | TTTCCAGGTGCCAAATGTAG | TATATTGTGACAAAAGAAAGGCAC | 55 |
| D12mit2* | chr12 | 22,650,702 | GTGGCTCTTTTCCTTAGGGA | TCGGCTTCTGAATGTATTGG | 55 |
| D12rat36 | chr12 | 37,521,884 | TGTAGAGAATCTAAAGCCCTCTTG | GAGGGACTTGGGAAGAGTGTT | 61 |
| D13rat60 | chr13 | 26,919,274 | ACGGCACAATCAATTGATGA | AAACTCACATGCTTGCACGT | 61 |
| LCA* | chr13 | 52,147,717 | TACAGAGCAAGCTCCAGGAC | TGTTTCTAATCCATAGGAAGTGC | 59 |
| D13rat131 | chr13 | 83,263,753 | CAGAATCTTGTGCCTCCTGA | TATCGACTTGCCATTGTTCG | 55 |
| D14rat72 | chr14 | 8,058,678 | CAGCAACACACACACCATACA | AGGAAGTCACACCACAAGGG | 61 |
| ALB* | chr14 | 19,191,541 | TTTTCGTAGTAACGGAAGCC | TAAGGATTCTCAGATGCAAATG | 59 |
| D14wox14 | chr14 | 22,009,849 | ACTTAATTACACACACAAACACAGA | CTTTGCTTTCTTTTAGCCATTT | 61 |
| D14rat39 | chr14 | 77,239,784 | GGCTGATCCCCAAGTGTCTA | TGAATCAAAAGGGGCATTTC | 59 |
| D15rat55 | chr15 | 211,024 | TATCCTCTTCTGCCCTCCAA | CCCACCCAGATGTGTTCTCT | 59 |
| D15rat123 | chr15 | 59,651,151 | CCCCTACATCCATGGCATAC | TTCCCCCTCTTGTTCTTCCT | 55 |
| D16wox9* | chr16 | 18,740,885 | ACCTACACGGACACATGGTG | GTTGTACTTCCTGATTTCCCTTT | 61 |
| D16rat15 | chr16 | 80,316,290 | ATCGGTAAACAAATCGCTGG | TGGCATCTTTCGCTTAGACA | 57 |
| D17rat78 | chr17 | 42,541,872 | TGCATGAGTCAATCTTCAATCA | CAGACTTCCTATTTTCCTATCTGCA | 57 |
| D17wox12* | chr17 | 63,994,211 | AGGAAATTAAGAGAAGTTGGGACT | TATGCTCTTTGGGCAGCTTA | 55 |
| D18wox7 | chr18 | 15,539,551 | GTGGAAAGCCTTCTGTTCAA | AGAATTCAATAATAACAGTCCCACT | 61 |
| TILP* | chr18 | 38,177,441 | AATCACTGGATGCTGGAAGA | AGGGAGCAGAACTACTAAAGATACA | 57 |
| D18rat116 | chr18 | 40,057,808 | TGCATATCCTTTTGCAACCA | TCCACTGTTCCAGGGAAGAC | 55 |
| D18rat8 | chr18 | 71,569,072 | GCGGCTGGAGTTATGACTTT | GCCCACTCCTGTGTGAGATT | 61 |
| D19wox8 | chr19 | 24,455,726 | TGCCCGTCTCTGTTACTCAT | CAAGAACCCTGAGGCAATAA | 59 |
| D19wox9 | chr19 | 42,099,630 | GTACAGGAACACTGCCCTTG | GGAATGTGTATGTGTGATGACG | 55 |
| AGT* | chr19 | 69,426,540 | TATGTAACTCAACGCCAGCC | GAAGCCCTAGTGGCAGATG | 59 |
| D20wox3* | chr20 | 4,855,475 | AGGAAATGGGTTTCAGTTCC | CAGGATTCTGTGGCAATCCG | 57 |
| D20mit4 | chr20 | 36,923,763 | TAAATTCAGAGTGTTGTTCCTGC | CACACCCAATGAATGCTCTG | 59 |
| DXwox6* | chrX | 28,435,436 | GTTTTCCCGCTTCACCAG | AGAAGGAGAAAGCGACCG | 59 |
| DXmgh9 | chrX | 70,352,120 | GTTGTCTGGTGCCAAAAGGT | AAGGATGATCTTAAGCTCCTGG | 59 |
表1 59个STR标记的引物序列及退火温度
Table 1 Primer sequences and annealing temperature of 59 STR markers
| 标记Marker | 染色体Chromosome | 物理位置/bp# Physical position/bp | 正向引物序列(5’→3‘)Upstream primer sequence(5’→3‘) | 反向引物序列(5’→3‘)Downstream primer sequence(5’→3‘) | 退火温度/℃ Annealing temperature/℃ |
|---|---|---|---|---|---|
| D1wox18 | chr1 | 100,133,276 | TCACTCTCATTAAACTAGGAATGC | ACTGTTGGGTAACAAAGTTATGG | 53 |
| D1rat345* | chr1 | 112,693,166 | TTCACACAAATGCCACCAGT | CAAAGAGATAGGGCGACAGG | 59 |
| D1wox21 | chr1 | 198,655,873 | GACATGGTGAGTGAAGACCC | TGAGGCTATCCTGAGCTAACTC | 59 |
| D1mgh14* | chr1 | 278,228,767 | CCGCACTGAGCTCTCAGAG | CCCAACCATTGAGCTAGTAAGG | 61 |
| D2rat3 | chr2 | 8,579,280 | TCTCTGGTGGATACCTCACAAA | ACATGATGTGGAATTCCCCC | 59 |
| D2mgh26* | chr2 | 105,075,219 | ATTGGTGATCAGATTTTTTTCTCC | CAGTTGAAAATTCCTGTATCTGG | 57 |
| D2wox15* | chr2 | 188,450,000 | GGTGCTAGTAGACAATAAGATAGAT | TTCATGAGTTTTCACTGTTTGC | 53 |
| D2Arb20 | chr2 | 200,585,161 | ACTAGAGTCAGGAGGCAAAACC | ACCCTATGTGAAATGGTGATGC | 59 |
| D3wox9* | chr3 | 51,821,710 | TTCAGGTATGATTCGGGAAC | ATTGGCGATATCATTTCACTAAC | 59 |
| RH94469 | chr3 | 93,380,189 | CTTATGTTACCTCACAGCCTGG | GGGTGGGCCATCTTTATAATC | 61 |
| D4wox27* | chr4 | 3,044,017 | GGCCTTCATTTGTCCTTTGA | CACCATTATGCCCAGCCTTA | 59 |
| D4Arb10* | chr4 | 61,708,051 | ACTCATCTTCTGAGGAGTAGCG | TTCACAGTCTATTTGGTAGGGC | 59 |
| D4mit15* | chr4 | 111,809,356 | CCTAATGACAGCAATAGAAACTACG | TTTTGGTACTGGAGAAAGTTGG | 61 |
| D5mit18 | chr5 | 17,062,312 | GCAAGACAGGCTTCCACAAT | AATACTGCCGGAGCAGAAAA | 59 |
| D5Hmgc2* | chr5 | 78,041,475 | ATGTTCCCTGATGTCCCTTC | AAACCATGCCACGTGAAC | 53 |
| D5wox11 | chr5 | 138,180,823 | CTTAGGTCAGCTGGTCCCTAG | CATGGTGCTTTTCCTTGTTG | 55 |
| D6rat105 | chr6 | 19,888,103 | GCATGAGGCAGAGCCAATTA | GCCAGTTTGCCTTCACTTTC | 59 |
| D6mit1* | chr6 | 99,273,921 | TTAGGAGAGAACTGAAAGTTGTCC | ATGGTGCACTATGGTGGTCA | 61 |
| D6wox12 | chr6 | 138,065,848 | CTTCCCTACCTTCTACAACACTAAG | TAAACCAAAGAGCATGATGAAG | 61 |
| D7wox14 | chr7 | 102,588,256 | GGGAGTAAAAGAGTGCATTCC | TACCCCAATCCTGAACCAC | 61 |
| D7wox24 | chr7 | 115,250,056 | AGCCTCAATCTGGCCTAGCTC | TGACTCTTGACAATGGCACA | 59 |
| D7mgh3* | chr7 | 123,632,617 | CGGCATGTGTCTCTGTGTG | TGACTTCTGGTGTCCTCACG | 59 |
| D8wox4* | chr8 | 50,529,563 | GATTTGAAGCGATTGTCCAT | GTCTAGCTGCCCACAGGAG | 57 |
| D8rat14* | chr8 | 108,068,142 | GCCCATACGTTGCATCAAGT | GGCCGGTCTAATTATTTCTTCA | 59 |
| D8wox11 | chr8 | 116,873,553 | AGTGGGAATGCTTAGCCAAA | CAGGCCTTTACACCTTCCTG | 57 |
| D9mit2* | chr9 | 71,771,288 | TGACCAGTTAGCCCTTTCCA | GGGAGCAGGGTTCTACACAG | 59 |
| D9rat110 | chr9 | 102,531,710 | TGGGTGCTTAATTTTGTTTGTTT | GCTCCTAACCCATCCAACTTT | 55 |
| D10rat199 | chr10 | 1,381,054 | TGCCAAATAATGATTGTTCCTTC | CCTCCTCTGGCTTCTTGAGA | 55 |
| D10wox12 | chr10 | 56,220,298 | CGCTGGGAGGAAAGAGAG | AGGCTTGCACTCTGTGTTTG | 59 |
| D10mit12* | chr10 | 94,240,320 | ATGGGAGGGAACAAGTCTTC | GAGGGAGAGAGAAAGAGAGACAG | 61 |
| D11wox3* | chr11 | 60,397,957 | CACTTTGAGGCCATTCTGAA | CCTTCTCTTTGTGAAAAATAAAGTC | 59 |
| D11mgh3* | chr11 | 69,649,708 | GGAGCTGAAATACGAGAGAAATAA | GTCCTGCTGGCTGTGCAT | 59 |
| D11wox2 | chr11 | 80,357,513 | TTTCCAGGTGCCAAATGTAG | TATATTGTGACAAAAGAAAGGCAC | 55 |
| D12mit2* | chr12 | 22,650,702 | GTGGCTCTTTTCCTTAGGGA | TCGGCTTCTGAATGTATTGG | 55 |
| D12rat36 | chr12 | 37,521,884 | TGTAGAGAATCTAAAGCCCTCTTG | GAGGGACTTGGGAAGAGTGTT | 61 |
| D13rat60 | chr13 | 26,919,274 | ACGGCACAATCAATTGATGA | AAACTCACATGCTTGCACGT | 61 |
| LCA* | chr13 | 52,147,717 | TACAGAGCAAGCTCCAGGAC | TGTTTCTAATCCATAGGAAGTGC | 59 |
| D13rat131 | chr13 | 83,263,753 | CAGAATCTTGTGCCTCCTGA | TATCGACTTGCCATTGTTCG | 55 |
| D14rat72 | chr14 | 8,058,678 | CAGCAACACACACACCATACA | AGGAAGTCACACCACAAGGG | 61 |
| ALB* | chr14 | 19,191,541 | TTTTCGTAGTAACGGAAGCC | TAAGGATTCTCAGATGCAAATG | 59 |
| D14wox14 | chr14 | 22,009,849 | ACTTAATTACACACACAAACACAGA | CTTTGCTTTCTTTTAGCCATTT | 61 |
| D14rat39 | chr14 | 77,239,784 | GGCTGATCCCCAAGTGTCTA | TGAATCAAAAGGGGCATTTC | 59 |
| D15rat55 | chr15 | 211,024 | TATCCTCTTCTGCCCTCCAA | CCCACCCAGATGTGTTCTCT | 59 |
| D15rat123 | chr15 | 59,651,151 | CCCCTACATCCATGGCATAC | TTCCCCCTCTTGTTCTTCCT | 55 |
| D16wox9* | chr16 | 18,740,885 | ACCTACACGGACACATGGTG | GTTGTACTTCCTGATTTCCCTTT | 61 |
| D16rat15 | chr16 | 80,316,290 | ATCGGTAAACAAATCGCTGG | TGGCATCTTTCGCTTAGACA | 57 |
| D17rat78 | chr17 | 42,541,872 | TGCATGAGTCAATCTTCAATCA | CAGACTTCCTATTTTCCTATCTGCA | 57 |
| D17wox12* | chr17 | 63,994,211 | AGGAAATTAAGAGAAGTTGGGACT | TATGCTCTTTGGGCAGCTTA | 55 |
| D18wox7 | chr18 | 15,539,551 | GTGGAAAGCCTTCTGTTCAA | AGAATTCAATAATAACAGTCCCACT | 61 |
| TILP* | chr18 | 38,177,441 | AATCACTGGATGCTGGAAGA | AGGGAGCAGAACTACTAAAGATACA | 57 |
| D18rat116 | chr18 | 40,057,808 | TGCATATCCTTTTGCAACCA | TCCACTGTTCCAGGGAAGAC | 55 |
| D18rat8 | chr18 | 71,569,072 | GCGGCTGGAGTTATGACTTT | GCCCACTCCTGTGTGAGATT | 61 |
| D19wox8 | chr19 | 24,455,726 | TGCCCGTCTCTGTTACTCAT | CAAGAACCCTGAGGCAATAA | 59 |
| D19wox9 | chr19 | 42,099,630 | GTACAGGAACACTGCCCTTG | GGAATGTGTATGTGTGATGACG | 55 |
| AGT* | chr19 | 69,426,540 | TATGTAACTCAACGCCAGCC | GAAGCCCTAGTGGCAGATG | 59 |
| D20wox3* | chr20 | 4,855,475 | AGGAAATGGGTTTCAGTTCC | CAGGATTCTGTGGCAATCCG | 57 |
| D20mit4 | chr20 | 36,923,763 | TAAATTCAGAGTGTTGTTCCTGC | CACACCCAATGAATGCTCTG | 59 |
| DXwox6* | chrX | 28,435,436 | GTTTTCCCGCTTCACCAG | AGAAGGAGAAAGCGACCG | 59 |
| DXmgh9 | chrX | 70,352,120 | GTTGTCTGGTGCCAAAAGGT | AAGGATGATCTTAAGCTCCTGG | 59 |
标记 Marker | 染色体 Chromosome | 物理位置/bp# Physical position /bp | 各品系的产物长度/bp Product size of each strain/bp | 多态性数量(Number of polymorphisms) | ||||
|---|---|---|---|---|---|---|---|---|
| F344 | BN | DA | Lewis | PVG | ||||
| D1wox18 | chr1 | 100,133,276 | 122 | 120 | 122 | 120 | 118 | 3 |
| D1rat345* | chr1 | 112,693,166 | 125 | 125 | 121 | 125 | 129 | 3 |
| D1wox21 | chr1 | 198,655,873 | 85 | 97 | 73 | 95 | 97 | 4 |
| D1mgh14* | chr1 | 278,228,767 | 142 | 120 | 150 | 150 | 140 | 4 |
| D2rat3 | chr2 | 8,579,280 | 126 | 122 | 124 | 117 | 122 | 4 |
| D2mgh26* | chr2 | 105,075,219 | 126 | 132 | 134 | 134 | 126 | 3 |
| D2wox15* | chr2 | 188,450,000 | 151 | 131 | 133 | 133 | 145 | 4 |
| D2Arb20 | chr2 | 200,585,161 | 231 | 235 | 207 | 215 | 207 | 4 |
| D3wox9* | chr3 | 51,821,710 | 117 | 123 | 115 | 121 | 121 | 4 |
| RH94469 | chr3 | 93,380,189 | 239 | 239 | 239 | 227 | 221 | 3 |
| D4wox27* | chr4 | 3,044,017 | 214 | 197 | 219 | 197 | 214 | 3 |
| D4Arb10* | chr4 | 61,708,051 | 389 | 392 | 389 | 389 | 409 | 3 |
| D4mit15* | chr4 | 111,809,356 | 215 | 215 | 215 | 215 | 202 | 2 |
| D5mit18 | chr5 | 17,062,312 | 235 | 219 | 225 | 235 | 231 | 4 |
| D5wox11 | chr5 | 138,180,823 | 137 | 123 | 137 | 137 | 137 | 2 |
| D6rat105 | chr6 | 19,888,103 | 229 | 225 | 204 | 227 | 227 | 4 |
| D6mit1* | chr6 | 99,273,921 | 134 | 122 | 112 | 134 | 134 | 3 |
| D6wox12 | chr6 | 138,065,848 | 127 | 127 | 135 | 127 | 127 | 2 |
| D7wox14 | chr7 | 102,588,256 | 120 | 126 | 122 | 118 | 124 | 5 |
| D7wox24 | chr7 | 115,250,056 | 133 | 133 | 133 | 133 | 129 | 2 |
| D7mgh3* | chr7 | 123,632,617 | 233 | 243 | 233 | 243 | 233 | 2 |
| D8wox4* | chr8 | 50,529,563 | 131 | 119 | 129 | 131 | 117 | 4 |
| D8rat14* | chr8 | 108,068,142 | 169 | 161 | 161 | 161 | 169 | 2 |
| D8wox11 | chr8 | 116,873,553 | 183 | 191 | 181 | 191 | 183 | 3 |
| D9mit2* | chr9 | 71,771,288 | 185 | 185 | 190 | 190 | 185 | 2 |
| D9rat110 | chr9 | 102,531,710 | 146 | 158 | 169 | 158 | 146 | 3 |
| D10rat199 | chr10 | 1,381,054 | 190 | 188 | 188 | 188 | 188 | 2 |
| D10wox12 | chr10 | 56,220,298 | 95 | 95 | 95 | 91 | 91 | 2 |
| D10mit12* | chr10 | 94,240,320 | 142 | 144 | 144 | 144 | 144 | 2 |
| D11wox3* | chr11 | 60,397,957 | 189 | 183 | 183 | 185 | 181 | 4 |
| D11mgh3* | chr11 | 69,649,708 | 150 | 166 | 186 | 145 | 145 | 4 |
| D11wox2 | chr11 | 80,357,513 | 152 | 146 | 148 | 146 | 148 | 3 |
| D12mit2* | chr12 | 22,650,702 | 187 | 215 | 187 | 206 | 196 | 4 |
| D12rat36 | chr12 | 37,521,884 | 205 | 202 | 205 | 188 | 190 | 4 |
| D13rat60 | chr13 | 26,919,274 | 131 | 125 | 125 | 127 | 127 | 3 |
| D13rat131 | chr13 | 83,263,753 | 149 | 151 | 149 | 134 | 134 | 3 |
| D14rat72 | chr14 | 8,058,678 | 152 | 181 | 162 | 179 | 152 | 4 |
| ALB* | chr14 | 19,191,541 | 136 | 120 | 136 | 133 | 136 | 3 |
| D14wox14 | chr14 | 22,009,849 | 122 | 112 | 122 | 108 | 108 | 3 |
| D14rat39 | chr14 | 77,239,784 | 240 | 248 | 240 | 248 | 240 | 2 |
| D15rat55 | chr 15 | 211,024 | 210 | 228 | 228 | 228 | 228 | 2 |
| D15rat123 | chr15 | 59,651,151 | 136 | 132 | 130 | 122 | 120 | 5 |
| D16wox9* | chr16 | 18,740,885 | 145 | 137 | 191 | 145 | 145 | 3 |
| D16rat15 | chr16 | 80,316,290 | 142 | 133 | 144 | 133 | 144 | 3 |
| D17rat78 | chr17 | 42,541,872 | 151 | 157 | 155 | 155 | 155 | 3 |
| D17wox12* | chr17 | 63,994,211 | 136 | 132 | 115 | 119 | 115 | 4 |
| D18wox7 | chr18 | 15,539,551 | 148 | 142 | 189 | 193 | 189 | 4 |
| TILP* | chr18 | 38,177,441 | 262 | 240 | 262 | 242 | 242 | 3 |
| D18rat116 | chr18 | 40,057,808 | 219 | 223 | 219 | 223 | 225 | 3 |
| D18rat8 | chr18 | 71,569,072 | 155 | 165 | 157 | 165 | 161 | 4 |
| D19wox8 | chr19 | 24,455,726 | 124 | 126 | 111 | 124 | 111 | 3 |
| D19wox9 | chr19 | 42,099,630 | 154 | 158 | 158 | 156 | 160 | 4 |
| D20wox3* | chr20 | 4,855,475 | 116 | 110 | 120 | 128 | 114 | 5 |
| D20mit4 | chr20 | 36,923,763 | 215 | 201 | 192 | 198 | 215 | 4 |
| Dxwox6* | chrX | 28,435,436 | 171 | 138 | 173 | 138 | 175 | 4 |
| DXmgh9 | chrX | 70,352,120 | 120 | 146 | 146 | 146 | 120 | 2 |
纯合度 homozygosity | / | / | 100% | 100% | 100% | 100% | 100% | / |
表2 56个STR标记在5个近交系大鼠中的分型结果
Table 2 Genotyping results of 56 STR markers in 5 inbred rat strains.
标记 Marker | 染色体 Chromosome | 物理位置/bp# Physical position /bp | 各品系的产物长度/bp Product size of each strain/bp | 多态性数量(Number of polymorphisms) | ||||
|---|---|---|---|---|---|---|---|---|
| F344 | BN | DA | Lewis | PVG | ||||
| D1wox18 | chr1 | 100,133,276 | 122 | 120 | 122 | 120 | 118 | 3 |
| D1rat345* | chr1 | 112,693,166 | 125 | 125 | 121 | 125 | 129 | 3 |
| D1wox21 | chr1 | 198,655,873 | 85 | 97 | 73 | 95 | 97 | 4 |
| D1mgh14* | chr1 | 278,228,767 | 142 | 120 | 150 | 150 | 140 | 4 |
| D2rat3 | chr2 | 8,579,280 | 126 | 122 | 124 | 117 | 122 | 4 |
| D2mgh26* | chr2 | 105,075,219 | 126 | 132 | 134 | 134 | 126 | 3 |
| D2wox15* | chr2 | 188,450,000 | 151 | 131 | 133 | 133 | 145 | 4 |
| D2Arb20 | chr2 | 200,585,161 | 231 | 235 | 207 | 215 | 207 | 4 |
| D3wox9* | chr3 | 51,821,710 | 117 | 123 | 115 | 121 | 121 | 4 |
| RH94469 | chr3 | 93,380,189 | 239 | 239 | 239 | 227 | 221 | 3 |
| D4wox27* | chr4 | 3,044,017 | 214 | 197 | 219 | 197 | 214 | 3 |
| D4Arb10* | chr4 | 61,708,051 | 389 | 392 | 389 | 389 | 409 | 3 |
| D4mit15* | chr4 | 111,809,356 | 215 | 215 | 215 | 215 | 202 | 2 |
| D5mit18 | chr5 | 17,062,312 | 235 | 219 | 225 | 235 | 231 | 4 |
| D5wox11 | chr5 | 138,180,823 | 137 | 123 | 137 | 137 | 137 | 2 |
| D6rat105 | chr6 | 19,888,103 | 229 | 225 | 204 | 227 | 227 | 4 |
| D6mit1* | chr6 | 99,273,921 | 134 | 122 | 112 | 134 | 134 | 3 |
| D6wox12 | chr6 | 138,065,848 | 127 | 127 | 135 | 127 | 127 | 2 |
| D7wox14 | chr7 | 102,588,256 | 120 | 126 | 122 | 118 | 124 | 5 |
| D7wox24 | chr7 | 115,250,056 | 133 | 133 | 133 | 133 | 129 | 2 |
| D7mgh3* | chr7 | 123,632,617 | 233 | 243 | 233 | 243 | 233 | 2 |
| D8wox4* | chr8 | 50,529,563 | 131 | 119 | 129 | 131 | 117 | 4 |
| D8rat14* | chr8 | 108,068,142 | 169 | 161 | 161 | 161 | 169 | 2 |
| D8wox11 | chr8 | 116,873,553 | 183 | 191 | 181 | 191 | 183 | 3 |
| D9mit2* | chr9 | 71,771,288 | 185 | 185 | 190 | 190 | 185 | 2 |
| D9rat110 | chr9 | 102,531,710 | 146 | 158 | 169 | 158 | 146 | 3 |
| D10rat199 | chr10 | 1,381,054 | 190 | 188 | 188 | 188 | 188 | 2 |
| D10wox12 | chr10 | 56,220,298 | 95 | 95 | 95 | 91 | 91 | 2 |
| D10mit12* | chr10 | 94,240,320 | 142 | 144 | 144 | 144 | 144 | 2 |
| D11wox3* | chr11 | 60,397,957 | 189 | 183 | 183 | 185 | 181 | 4 |
| D11mgh3* | chr11 | 69,649,708 | 150 | 166 | 186 | 145 | 145 | 4 |
| D11wox2 | chr11 | 80,357,513 | 152 | 146 | 148 | 146 | 148 | 3 |
| D12mit2* | chr12 | 22,650,702 | 187 | 215 | 187 | 206 | 196 | 4 |
| D12rat36 | chr12 | 37,521,884 | 205 | 202 | 205 | 188 | 190 | 4 |
| D13rat60 | chr13 | 26,919,274 | 131 | 125 | 125 | 127 | 127 | 3 |
| D13rat131 | chr13 | 83,263,753 | 149 | 151 | 149 | 134 | 134 | 3 |
| D14rat72 | chr14 | 8,058,678 | 152 | 181 | 162 | 179 | 152 | 4 |
| ALB* | chr14 | 19,191,541 | 136 | 120 | 136 | 133 | 136 | 3 |
| D14wox14 | chr14 | 22,009,849 | 122 | 112 | 122 | 108 | 108 | 3 |
| D14rat39 | chr14 | 77,239,784 | 240 | 248 | 240 | 248 | 240 | 2 |
| D15rat55 | chr 15 | 211,024 | 210 | 228 | 228 | 228 | 228 | 2 |
| D15rat123 | chr15 | 59,651,151 | 136 | 132 | 130 | 122 | 120 | 5 |
| D16wox9* | chr16 | 18,740,885 | 145 | 137 | 191 | 145 | 145 | 3 |
| D16rat15 | chr16 | 80,316,290 | 142 | 133 | 144 | 133 | 144 | 3 |
| D17rat78 | chr17 | 42,541,872 | 151 | 157 | 155 | 155 | 155 | 3 |
| D17wox12* | chr17 | 63,994,211 | 136 | 132 | 115 | 119 | 115 | 4 |
| D18wox7 | chr18 | 15,539,551 | 148 | 142 | 189 | 193 | 189 | 4 |
| TILP* | chr18 | 38,177,441 | 262 | 240 | 262 | 242 | 242 | 3 |
| D18rat116 | chr18 | 40,057,808 | 219 | 223 | 219 | 223 | 225 | 3 |
| D18rat8 | chr18 | 71,569,072 | 155 | 165 | 157 | 165 | 161 | 4 |
| D19wox8 | chr19 | 24,455,726 | 124 | 126 | 111 | 124 | 111 | 3 |
| D19wox9 | chr19 | 42,099,630 | 154 | 158 | 158 | 156 | 160 | 4 |
| D20wox3* | chr20 | 4,855,475 | 116 | 110 | 120 | 128 | 114 | 5 |
| D20mit4 | chr20 | 36,923,763 | 215 | 201 | 192 | 198 | 215 | 4 |
| Dxwox6* | chrX | 28,435,436 | 171 | 138 | 173 | 138 | 175 | 4 |
| DXmgh9 | chrX | 70,352,120 | 120 | 146 | 146 | 146 | 120 | 2 |
纯合度 homozygosity | / | / | 100% | 100% | 100% | 100% | 100% | / |
大鼠品系 Strains | F344 | BN | DA | Lewis | PVG |
|---|---|---|---|---|---|
| F344 | - | 49 | 40 | 45 | 40 |
| BN | 1.775 | - | 44 | 35 | 49 |
| DA | 1.133 | 1.369 | - | 42 | 42 |
| Lewis | 1.438 | 0.899 | 1.244 | - | 40 |
| PVG | 1.133 | 1.680 | 1.244 | 1.133 | - |
表3 5个近交系大鼠两两品系间的STR标记等位基因总差异数及遗传距离
Table 3 Total number of STR markers allele differences and genetic distances between pairwise inbred rat strains
大鼠品系 Strains | F344 | BN | DA | Lewis | PVG |
|---|---|---|---|---|---|
| F344 | - | 49 | 40 | 45 | 40 |
| BN | 1.775 | - | 44 | 35 | 49 |
| DA | 1.133 | 1.369 | - | 42 | 42 |
| Lewis | 1.438 | 0.899 | 1.244 | - | 40 |
| PVG | 1.133 | 1.680 | 1.244 | 1.133 | - |
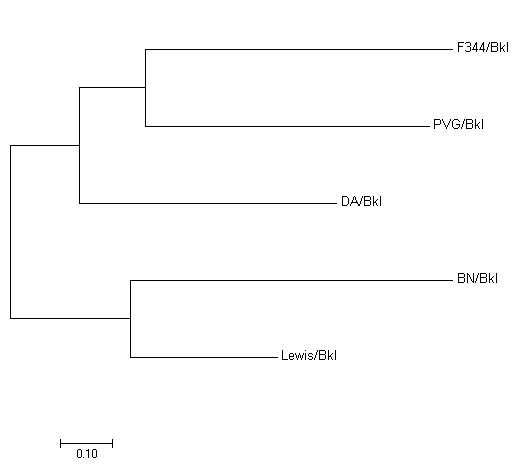
图1 5个近交系大鼠的NJ法遗传进化树
Figure 1 Neighbor-Joining (NJ) phylogenetic tree of 5 inbred rat strains.
染色体 Chromosome | STR标记组合 STR marker combinations | 染色体 Chromosome | STR标记组合 STR marker combinations |
|---|---|---|---|
| chr1 | D1wox18#、D1wox21&、D1mgh14& | chr11 | D11wox3&、D11wox2# |
| chr2 | D2rat3&、D2mgh26#、D2Arb20& | chr12 | D12mit2&、D12rat36& |
| chr3 | D3wox9&、RH94469# | chr13 | D13rat60#、D13rat131# |
| chr4 | D4wox27#、D4Arb10# | chr14 | D14rat72&、D14wox14# |
| chr5 | D5mit18&、D5wox11* | chr15 | D15rat55*、D15rat123† |
| chr6 | D6rat105&、D6mit1# | chr16 | D16wox9#、D16rat15# |
| chr7 | D7wox14†、D7mgh3* | chr17 | D17rat78#、D17wox12& |
| chr8 | D8wox4&、D8wox11# | chr18 | D18wox7&、D18rat8& |
| chr9 | D9mit2*、D9rat110# | chr19 | D19wox8#、D19wox9& |
| chr10 | D10rat199*、D10mit12* | chr20 | D20wox3†、D20mit4& |
表4 每条染色体的STR标记组合
Table 4 STR marker combinations for each chromosome
染色体 Chromosome | STR标记组合 STR marker combinations | 染色体 Chromosome | STR标记组合 STR marker combinations |
|---|---|---|---|
| chr1 | D1wox18#、D1wox21&、D1mgh14& | chr11 | D11wox3&、D11wox2# |
| chr2 | D2rat3&、D2mgh26#、D2Arb20& | chr12 | D12mit2&、D12rat36& |
| chr3 | D3wox9&、RH94469# | chr13 | D13rat60#、D13rat131# |
| chr4 | D4wox27#、D4Arb10# | chr14 | D14rat72&、D14wox14# |
| chr5 | D5mit18&、D5wox11* | chr15 | D15rat55*、D15rat123† |
| chr6 | D6rat105&、D6mit1# | chr16 | D16wox9#、D16rat15# |
| chr7 | D7wox14†、D7mgh3* | chr17 | D17rat78#、D17wox12& |
| chr8 | D8wox4&、D8wox11# | chr18 | D18wox7&、D18rat8& |
| chr9 | D9mit2*、D9rat110# | chr19 | D19wox8#、D19wox9& |
| chr10 | D10rat199*、D10mit12* | chr20 | D20wox3†、D20mit4& |
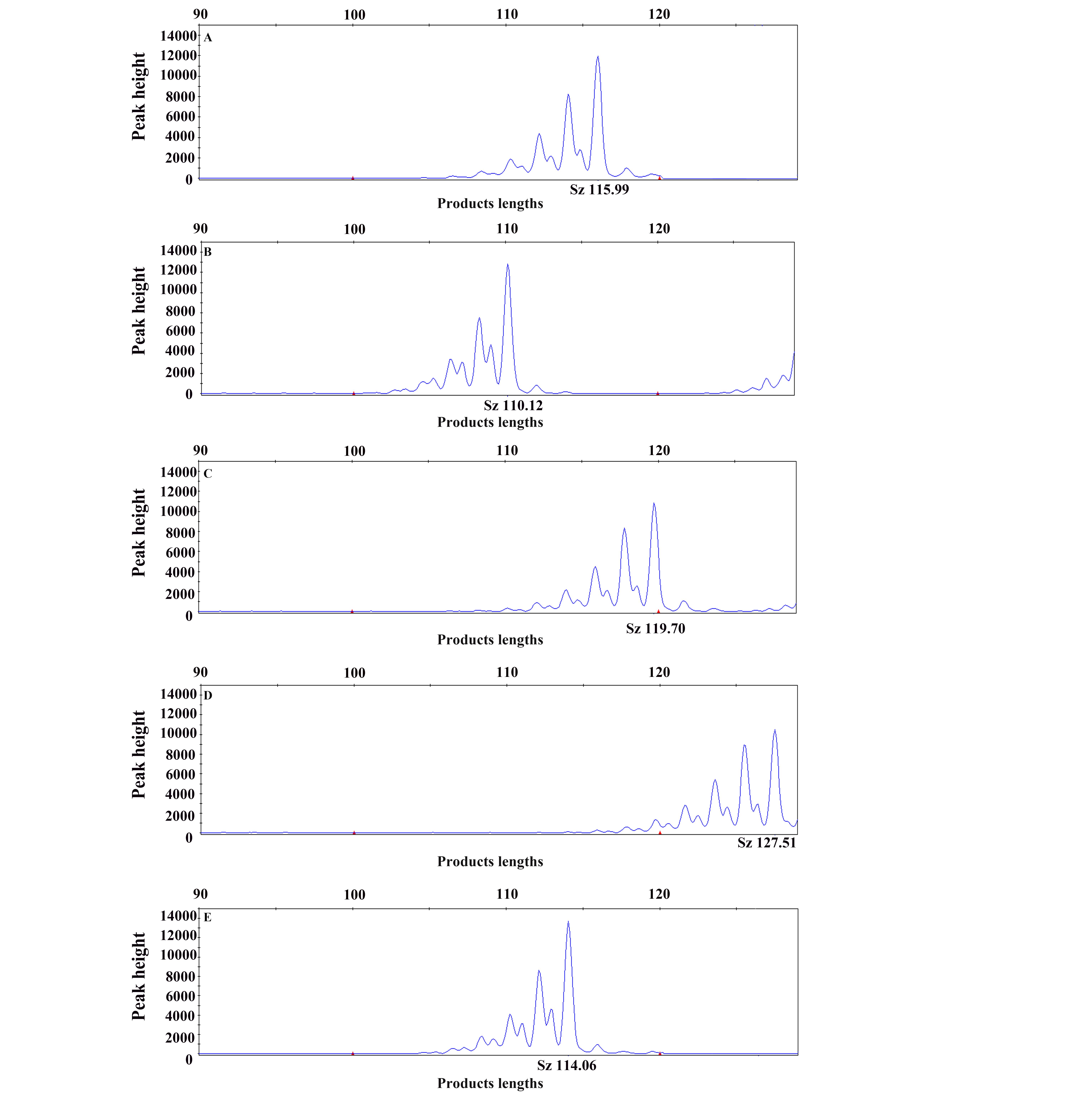
图2 D20wox3在5个近交系大鼠中的毛细管电泳峰图
Figure 2 Electropherogram of D20wox3 in 5 inbred rat strains
| [1] | GIBBS R A, WEINSTOCK G M, METZKER M L, et al. Genome sequence of the Brown Norway rat yields insights into mammalian evolution[J]. Nature, 2004, 428(6982): 493-521. DOI: 10.1038/nature02426 . |
| [2] | CARTER C S, RICHARDSON A, HUFFMAN D M, et al. Bring back the rat![J]. J Gerontol Ser A, 2020, 75(3): 405-415. DOI: 10.1093/gerona/glz298 . |
| [3] | JEWETT M, JIMENEZ-FERRER I, SWANBERG M. Astrocytic expression of GSTA4 is associated to dopaminergic neuroprotection in a rat 6-OHDA model of Parkinson's disease[J]. Brain Sci, 2017, 7(7): 73. DOI: 10.3390/brainsci7070073 . |
| [4] | NAKANO T, CHEN I H, GOTO S, et al. Hepatic miR-301a as a liver transplant rejection biomarker? and its role for interleukin-6 production in hepatocytes[J]. OMICS, 2017, 21(1): 55-66. DOI: 10.1089/omi.2016.0164 . |
| [5] | CHEN Z H, WANG C, WEI F X, et al. Adenovirus-mediated OX40Ig gene transfer induces long-term survival of orthotopic liver allograft in rats[J]. Transpl Immunol, 2018, 48: 32-38. DOI: 10.1016/j.trim.2018.02.010 . |
| [6] | ZAKRZEWICZ A, KUTLIMURATOV K, ATANASOVA S, et al. Attenuation of acute rat renal allograft rejection by apolipoprotein E-mimetic Peptide[J]. Transplantation, 2015, 99(5): 925-934. DOI: 10.1097/TP.0000000000000569 . |
| [7] | 国家市场监督管理总局, 国家标准化管理委员会. 实验动物 遗传质量控制: [S]. 北京: 中国标准出版社, 2022. |
| State Administration for Market Regulation, National Standardization Administration. Laboratory animal—Genetic quality control: [S]. Beijing: Standards Press of China, 2022. | |
| [8] | 李永明, 夏长友, 王海霞, 等. 近交系PVG大鼠的主要生物学特性的研究[J]. 畜牧兽医科技信息, 2008(4): 52-53. DOI: 10.3969/j.issn.1671-6027.2008.04.040 . |
| LI Y M, XIA C Y, WANG H X, et al. Study on the main biological characteristics of inbred PVG rats[J]. Chin J Anim Husb Vet Med, 2008(4): 52-53. DOI: 10.3969/j.issn.1671-6027.2008.04.040 . | |
| [9] | GÜNTHER M, NIMER F AL, PIEHL F, et al. Susceptibility to oxidative stress is determined by genetic background in neuronal cell cultures[J]. eNeuro, 2018, 5(2): ENEURO.0335-17.2018. DOI: 10.1523/ENEURO.0335-17.2018 . |
| [10] | BRYDA E C, RILEY L K. Multiplex microsatellite marker panels for genetic monitoring of common rat strains[J]. J Am Assoc Lab Anim Sci, 2008, 47(3): 37-41. |
| [11] | BENAVIDES F, RÜLICKE T, PRINS J B, et al. Genetic quality assurance and genetic monitoring of laboratory mice and rats: FELASA Working Group Report[J]. Lab Anim, 2020, 54(2): 135-148. DOI: 10.1177/0023677219867719 . |
| [12] | 李瑞生, 陈振文, 宋德光, 等. PCR扩增近交系大鼠微卫星位点DNA多态性的研究[J]. 遗传, 2001, 23(6): 539-543. DOI: 10.16288/j.yczz.2001.06.009 . |
| LI R S, CHEN Z W, SONG D G, et al. Rat gene polymorphism using PCR-analyzed microsatellites[J]. Hereditas (Beijing), 2001, 23(6): 539-543. DOI: 10.16288/j.yczz.2001.06.009 . | |
| [13] | 王越甲, 杨炜峰, 车路平, 等. 利用优选的微卫星标记建立常用大小鼠品系的遗传监测体系[J]. 实验动物科学, 2014, 31(3): 5-9, 14. DOI: 10.3969/j.issn.1006-6179.2014.03.002 . |
| WANG Y J, YANG W F, CHE L P, et al. Establishment of the genetic monitoring system in the mouse and rat by using selected microsatellite markers[J]. Lab Anim Sci, 2014, 31(3): 5-9, 14. DOI: 10.3969/j.issn.1006-6179.2014.03.002 . | |
| [14] | 郑龙, 李建辉, 王俊霞, 等. 微卫星DNA标记在近交系大鼠HFJ和MIJ遗传监测中的应用[J]. 中国实验动物学报, 2012, 20(2): 32-36. |
| ZHENG L, LI J H, WANG J X, et al. Microsatellite DNA analysis for genetic monitoring of inbred strain HFJ and MIJ rats[J]. Acta Lab Animalis Sci Sin, 2012, 20(2): 32-36. | |
| [15] | SERIKAWA T, KURAMOTO T, HILBERT P, et al. Rat gene mapping using PCR-analyzed microsatellites[J]. Genetics, 1992, 131(3): 701-721. DOI: 10.1093/genetics/131.3.701 . |
| [16] | KUMAR S, STECHER G, TAMURA K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets[J]. Mol Biol Evol, 2016, 33(7): 1870-1874. DOI: 10.1093/molbev/msw054 . |
| [17] | SAITOU N, NEI M. The neighbor-joining method: a new method for reconstructing phylogenetic trees[J]. Mol Biol Evol, 1987, 4(4): 406-425. DOI: 10.1093/oxfordjournals.molbev.a040454 . |
| [18] | FAHEY J R, KATOH H, MALCOLM R, et al. The case for genetic monitoring of mice and rats used in biomedical research[J]. Mamm Genome, 2013, 24(3-4): 89-94. DOI: 10.1007/s00335-012-9444-9 . |
| [19] | MASHIMO T, VOIGT B, TSURUMI T, et al. A set of highly informative rat simple sequence length polymorphism (SSLP) markers and genetically defined rat strains[J]. BMC Genet, 2006, 7: 19. DOI: 10.1186/1471-2156-7-19 . |
| [20] | KIM H, YOSHIHARA M, SUYAMA M. Comparative genomic analysis of inbred rat strains reveals the existence of ancestral polymorphisms[J]. Mamm Genome, 2020, 31(3-4): 86-94. DOI: 10.1007/s00335-020-09831-7 . |
| [21] | 林彦, 唐小玲, 王洪, 等. 近交系DA大鼠遗传学质量[J]. 中国比较医学杂志, 2009, 19(5): 59-61, 89. DOI: 10.3969/j.issn.1671-7856.2009.05.016 . |
| LIN Y, TANG X L, WANG H, et al. Detection of genes in the new inbred strain DA rat[J]. Chin J Comp Med, 2009, 19(5): 59-61, 89. DOI: 10.3969/j.issn.1671-7856.2009.05.016 . | |
| [22] | 朱婉月, 王洪, 刘巍, 等. PVG大鼠生化标记基因的测定[J]. 实验动物科学, 2024, 41(1): 29-32. DOI: 10.3969/j.issn.1006-6179.2024.01.006 . |
| ZHU W Y, WANG H, LIU W, et al. Determination of biochemical marker genes in PVG rats[J]. Lab Anim Sci, 2024, 41(1): 29-32. DOI: 10.3969/j.issn.1006-6179.2024.01.006 . | |
| [23] | 国家质量监督检验检疫总局, 国家标准化管理委员会. 实验动物 近交系小鼠、大鼠生化标记检测法: [S]. 北京: 中国标准出版社, 2009. |
| General Administration of Quality Supervision Inspection and Quarantine, National Standardization Administration. Laboratory animals—Methods for biochemical markers of inbred mice and rats: [S]. Beijing: Standards Press of China, 2009. |
| [1] | 孔志豪, 魏晓锋, 于灵芝, 冯丽萍, 朱琦, 施国君, 王晨. 鳞状皮屑裸小鼠木糖葡萄球菌的分离鉴定[J]. 实验动物与比较医学, 2025, 45(3): 368-375. |
| [2] | 赵丽亚, 倪丽菊, 张彩勤, 汤建平, 姚养正, 聂艳艳, 顾晓雪, 赵莹. 基于多重PCR-LDR技术建立近交系大鼠单核苷酸多态性遗传检测方案[J]. 实验动物与比较医学, 2023, 43(5): 548-558. |
| [3] | 巩淼淼, 周丽桂, 赵菊梅, 师长宏. 胃癌类器官在精准医学研究中的应用[J]. 实验动物与比较医学, 2022, 42(3): 248-254. |
| [4] | 成大欣, 徐丽然, 于清清, 高守翠, 王晓靖, 刘懿, 刘恩岐, 赵四海. 一种简单的gRNA体外筛选方法[J]. 实验动物与比较医学, 2018, 38(2): 126-129. |
| [5] | 李银银, 陈振文, 刘丽均, 李长龙, 刘欣, 吕建祎, 郭萌, 杜小燕. 单核苷酸多态性位点法对国内近交系小鼠遗传检测[J]. 实验动物与比较医学, 2018, 38(1): 16-21. |
| [6] | 周建华, 李志雄, 杨燕燕, 谢金东, 俞春英, 王训立. 应用微卫星DNA标记对福建亚种猕猴遗传多样性的分析[J]. 实验动物与比较医学, 2017, 37(6): 434-441. |
| [7] | 林丽芳, 李莉, 肖邦, 程继帅, 杨文静, 丛薇, 赵善民, 汤球, 崔淑芳. 应用通用荧光引物筛选裸鼹鼠微卫星位点方法的建立[J]. 实验动物与比较医学, 2016, 36(1): 76-80. |
| [8] | 林丽芳, 王洪, 赵善民, 肖邦, 孙智远, 张璐, 袁子彦, 于琛琳, 蔡丽萍, 徐晨, 汤球, 孙伟, 崔淑芳. 裸鼹鼠生化位点的初步研究[J]. 实验动物与比较医学, 2013, 33(6): 477-480. |
| [9] | 李爱学, 曾林, 尚世臣, 王鹏, 姚晓兰, 孙兆增. 黑线仓鼠白化突变系抑制差减文库的构建及初步分析[J]. 实验动物与比较医学, 2013, 33(1): 19-22. |
| [10] | 左宝芬, 杜小燕, 霍学云, 李振坤, 陈振文. 五个C57BL/6J小鼠生产群的微卫星检测及遗传学质量分析[J]. 实验动物与比较医学, 2012, 32(1): 51-55. |
| [11] | 王伟1,张传山2,郭毅3,谷瑞环3,丁雷4,姚刚1,陈学进3. 成体兔次黄嘌呤-鸟嘌呤磷酸核糖转移酶基因的打靶研究[J]. 实验动物与比较医学, 2010, 30(4): 233-240. |
| [12] | 余佳,蔡月琴,肖慧,陈民利. 实验兔RAPD反应体系的优化及引物筛选[J]. 实验动物与比较医学, 2009, 29(5): 341-346. |
| [13] | 王思锋,王希敏,侯海荣,刘可春,韩利文,陈锡强,王雪. 有机助溶剂对血管生成斑马鱼模型的适用性研究[J]. 实验动物与比较医学, 2008, 28(4): 238-242. |
| [14] | 吴海燕1,2,姚明2,闫明霞2,刘蕾2,孔韩卫2,薛整风1. 黑色素瘤小鼠高转移模型体内筛选和细胞的建立[J]. 实验动物与比较医学, 2008, 28(2): 67-73. |
| [15] | 张杰1, 2, 姚明2, 闫明霞2, 徐冰2, 刘蕾2, 孔韩卫2, 罗玉柱1. 小鼠结肠癌高转移模型体内筛选和建立[J]. 实验动物与比较医学, 2006, 26(3): 134-139. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||